RADIA: Radiology + AI A DICOM Viewer with AI on the Edge
How we built a complete medical imaging platform — DICOM viewer, 3D reconstruction via ray marching, multiplanar MPR and multi-model AI analysis — with zero servers, all running on Cloudflare Workers.
Traditional medical imaging tools are heavy desktop software, expensive licenses and workflows that haven't changed in 20 years. A radiologist opens a PACS viewer, manually reviews slice by slice, dictates their report and sends it to the clinician. What if that entire workflow could live in a browser, with AI analyzing every slice in parallel, generating findings with coordinates and severity, and enabling real-time collaboration?
That's RADIA. Not a prototype or a demo. It's a production platform that processes real DICOM — from dental CBCTs with 300+ slices to mammograms, CT scans and clinical dermatology photography — with three AI models working in consensus.
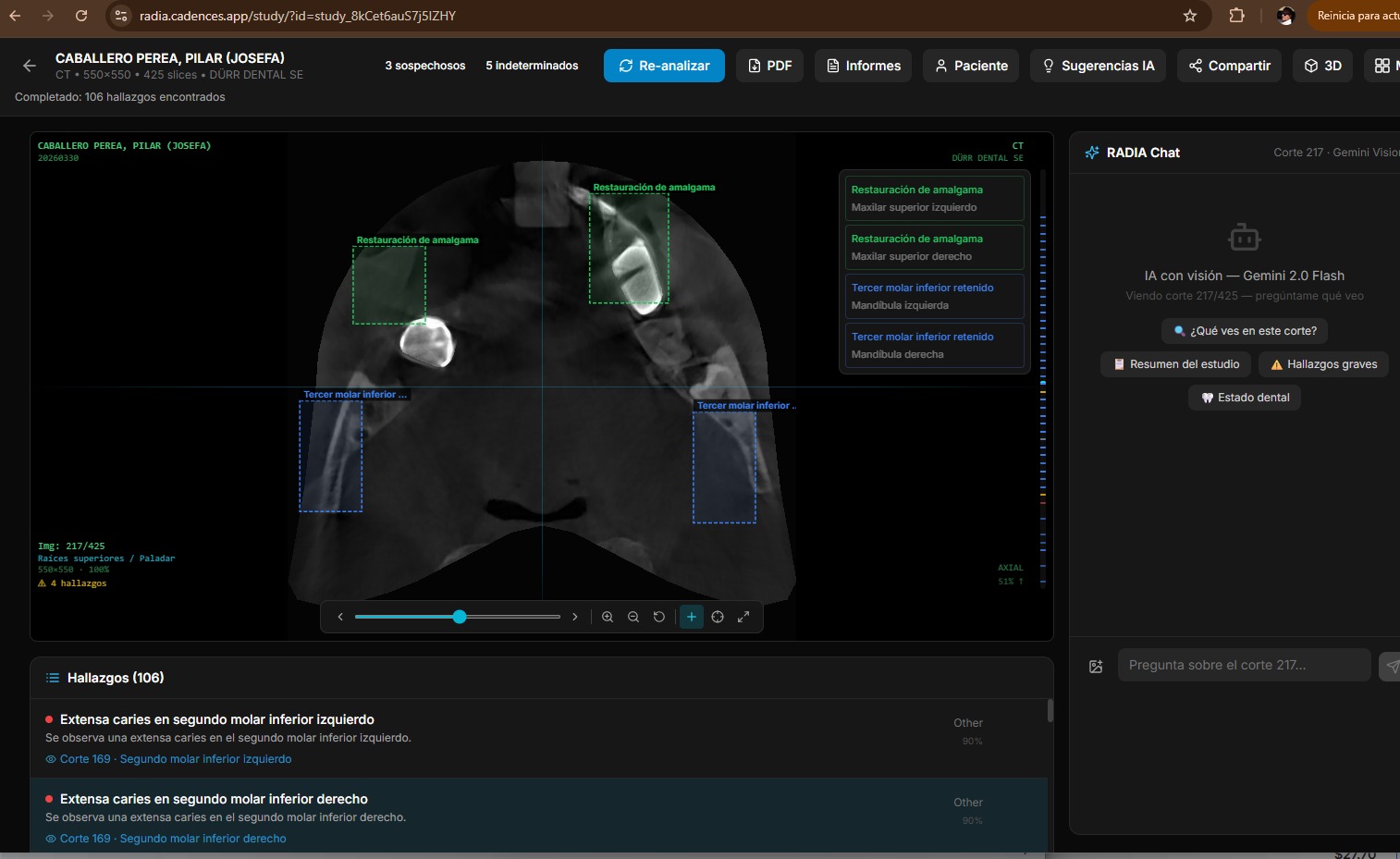
16 modalities, 4 models, 0 servers
RADIA supports 16 study types across 6 medical sectors, analyzed by Gemini 2.5 Flash + MedGemma 4B-IT + Llama 4 Scout + DeepSeek Reasoner, deployed entirely on Cloudflare Pages + Workers + D1 + R2. No EC2, no Kubernetes, no GPU bills.
Why current PACS doesn't scale
The conventional radiological workflow has three fundamental problems that have remained unsolved for decades:
Captive software
PACS viewers require local installation, per-seat licenses and specific hardware. A radiologist can't review a study from their phone on the way to the hospital.
Manual review
A dental CBCT has 200-400 slices. The doctor reviews them one by one looking for pathologies. A subtle finding on slice 247 can be missed due to visual fatigue.
Impossible collaboration
Sharing a study between professionals means CDs, USBs or hospital portals with VPN. There's no simple way to request a second opinion on a finding.
Edge-first: everything on Cloudflare
The most important decision was not to spin up servers. RADIA runs 100% on Cloudflare's edge: pages are static (Astro SSG), business logic runs on Workers, the database is D1 (distributed SQLite) and DICOM files live on R2 (S3-compatible). The result: 3-second deployments, global latency < 50ms, and zero infrastructure maintenance.
Astro 4 + React
SSG → static HTML with interactive React islands
Workers
16 API endpoints with JWT auth, AI analysis and DICOM proxy
D1
Distributed SQLite: users, studies, findings, chat, jobs
R2
DICOM file and PNG thumbnail storage
Client-side DICOM parsing with WASM
DICOM is the universal medical imaging standard, but it's a complex format: binary headers with nested tags, varied encodings (Little Endian, Big Endian, encapsulated) and JPEG 2000 compression that browsers don't natively understand.
Instead of parsing on the server (which would consume Worker resources), RADIA does all parsing on the client. The dicom-parser library extracts headers, and for JPEG 2000 pixels we use the OpenJPEG codec compiled to WebAssembly. The result: the Worker never touches pixel data, it only orchestrates.
ZIP (drag & drop)
→ JSZip extracts files
→ dicom-parser reads headers (patient, modality, geometry)
→ OpenJPEG WASM decodes JPEG 2000 → PNG thumbnails
→ POST /api/studies/upload (metadata)
→ PUT /api/dicom/{key} × N (binaries in parallel, 5 concurrent)
→ POST /api/studies/{id} action=finalize
→ Redirect → viewer with study readyThe trick is the dual storage layer: original DICOMs are stored in R2 for AI analysis, and in parallel we generate PNG thumbnails that are 10-50× smaller for fast navigation. The viewer preloads ±5 slices in each direction using the browser's Cache API, achieving an instant scroll experience even with 400-slice studies.
Progressive HQ Rendering — Native 16-bit
When the user stops scrolling and pauses on a slice for 800ms, RADIA downloads the original DICOM from R2, parses it in the browser (full 16-bit) and re-renders with the sensor's native dynamic range. An HD 16-bit badge appears in the PACS overlay.
While scrolling: instant 8-bit PNG (~50ms). When paused: 16-bit DICOM with original W/L (~200-500ms). Best of both worlds: fluidity + diagnostic quality.
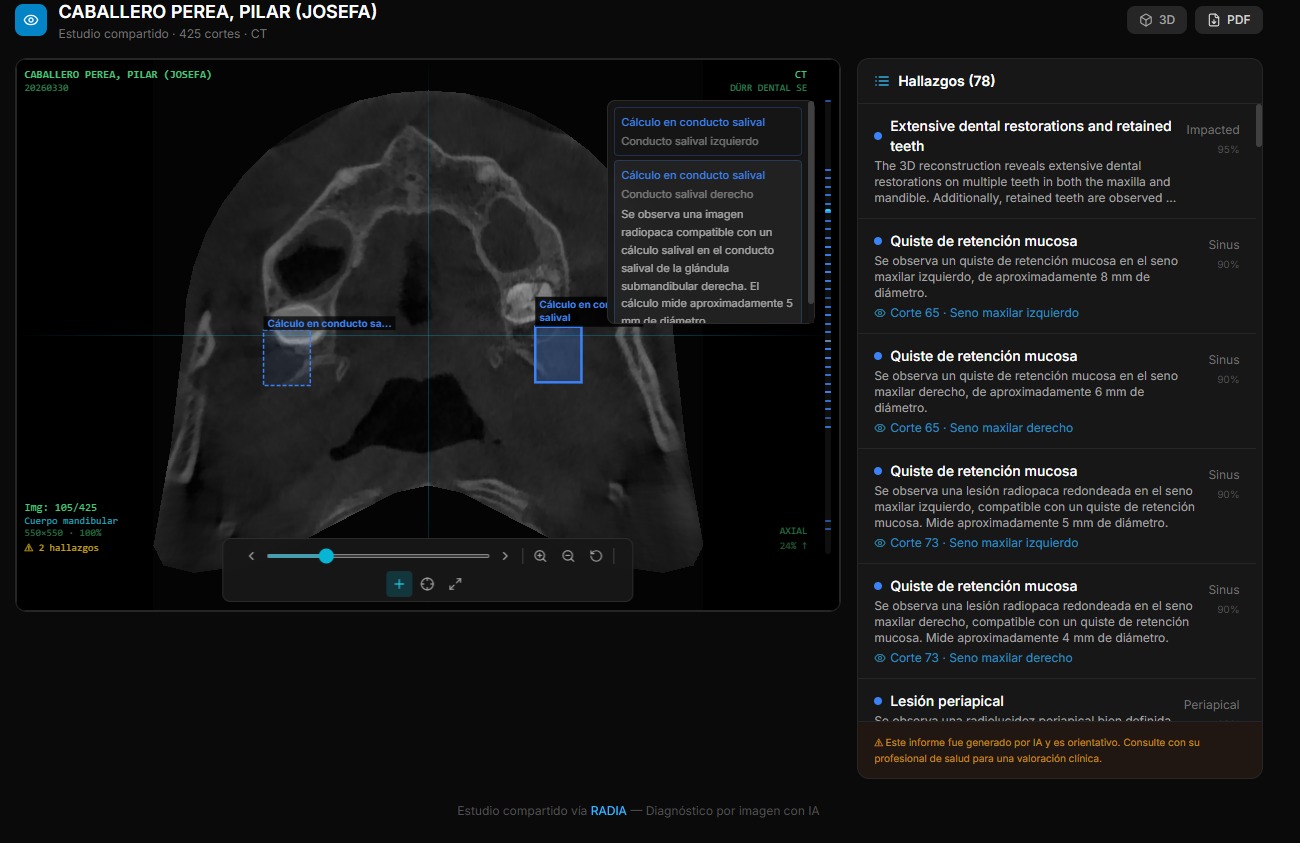
Volume rendering with raw WebGL2
For 3D reconstruction of volumetric images (dental CBCT, CT), we wrote a ray marcher from scratch in GLSL. No Three.js (400KB+), no 3D dependencies. Just two shaders, a 3D texture and inline 4×4 matrix math.
How ray marching works
PNGs from each slice are stacked into a WebGL2 3D texture (256³ on mobile, 768³ on desktop)
For each screen pixel, the fragment shader fires a ray through the volume
The ray marches in 160 steps (mobile) or 384 steps (desktop), sampling tissue density
A transfer function maps density → color: bone beige, soft tissue translucent, air invisible
Three render modes: Bone (volumetric compositing), MIP (maximum intensity projection), X-Ray (inverse projection)
The volume is cached in IndexedDB with a 7-day TTL, so reopening a study shows 3D instantly without rebuilding. On mobile, we reduce texture resolution and ray marching steps to maintain 30+ FPS with smooth touch rotation.
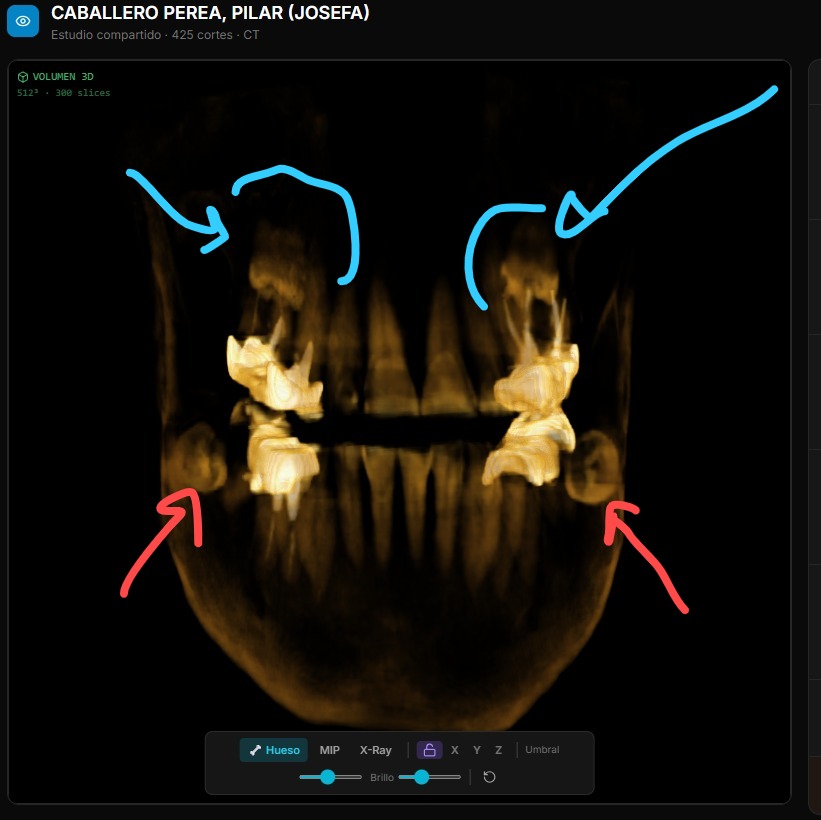
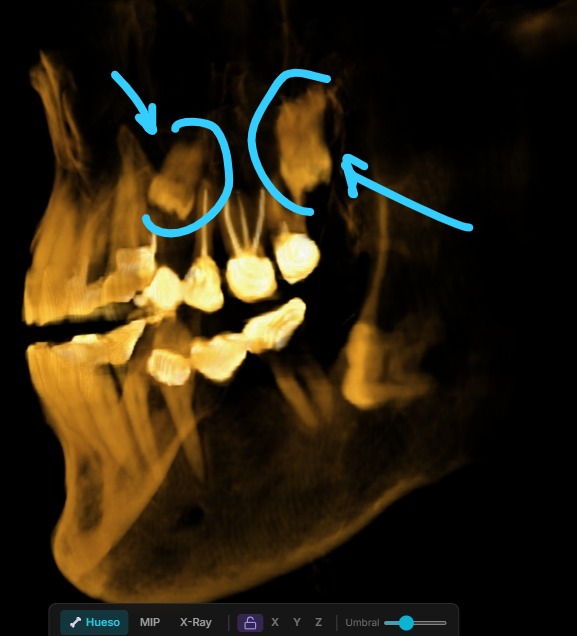
MPR: Multi-Planar Reconstruction
Beyond 3D, RADIA offers MPR view in a 2×2 grid — axial, coronal, sagittal plus a mini-3D — with synchronized crosshairs across all four views. Click any panel to expand it full screen. Everything recalculated client-side from the same WebGL2 3D texture.
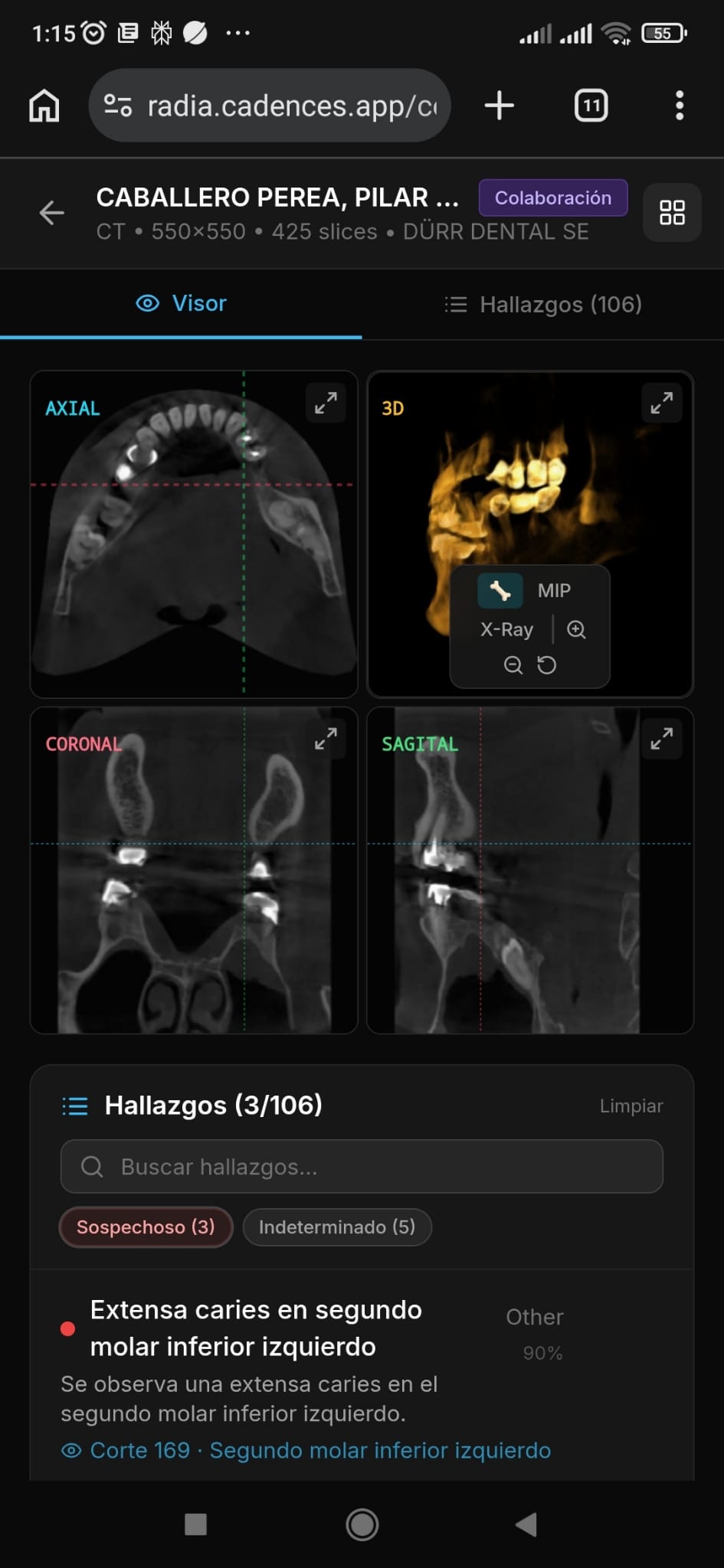
5 analysis modes, 4 models in cascade
RADIA doesn't have a single "analyze" endpoint. It has five distinct analysis modes, each designed for a different moment in the doctor's workflow. Before running any analysis, a pre-analysis dialog lets the doctor choose whether to complement existing findings, critically review them, or wipe and re-analyze from scratch — with an Asteroid Mode toggle and a reason field to guide the AI's focus.
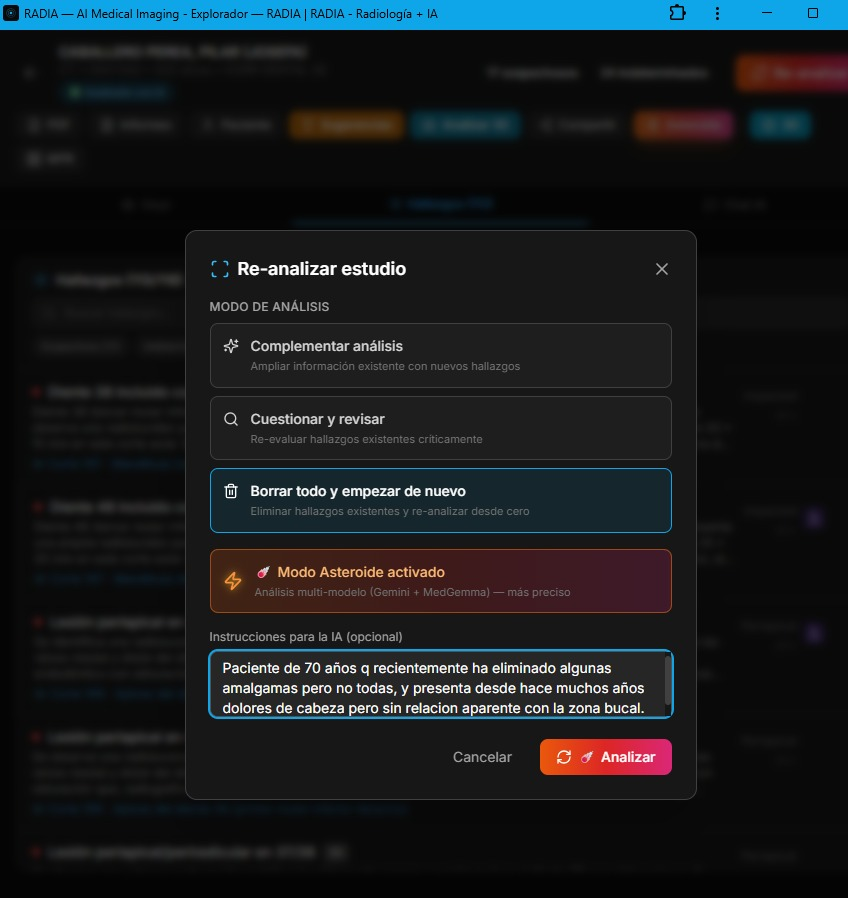
Scan 360°
Automated full-study analysis in batches. The frontend orchestrates batches to stay within the Workers timeout (30s). Samples every 8 slices, generating findings with title, severity, confidence, normalized bounding box and anatomical location.
Deep Analysis
Deep dive into a specific finding. Reads ±10 slices around the finding, generates clinical differential, clinical significance, recommended actions and confidence review.
Point-Ask
The doctor clicks anywhere on the image and asks "what do you see here?". The normalized coordinates (x, y) are sent along with the slice PNG. MedGemma gets extra weight here due to its medical specialization.
Viewport Capture
Captures the current 3D or MPR view as PNG, saves it to R2 and analyzes it. Useful for documenting volumetric findings that don't exist in a single 2D slice.
Treatment Suggestions
Synthesizes all study findings into a prioritized treatment plan. Takes into account severity, anatomical location, previous deep analyses and relationships between findings.
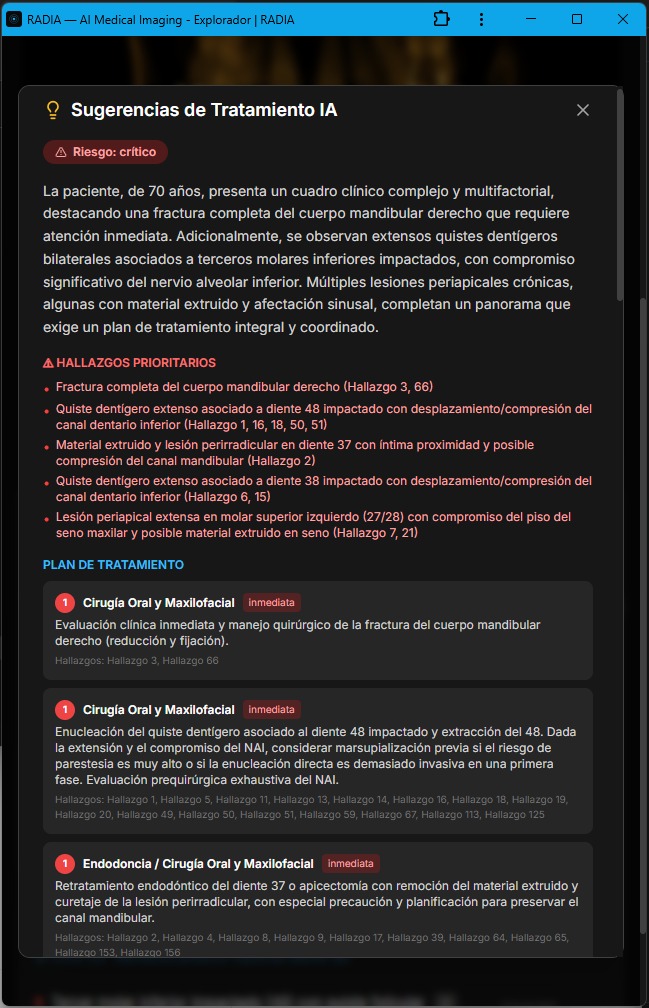
AI-generated treatment suggestions: risk level, priority findings and staged plan with urgency and referrals.
The four models are chained in priority cascade:
| Priority | Model | Provider | Role |
|---|---|---|---|
| 1 | Gemini 2.5 Flash | Google AI | Primary — multimodal vision with long context |
| 2 | Llama 4 Scout 17B | Groq | Fallback — if Gemini fails or is overloaded |
| 3 | MedGemma 4B-IT | Vertex AI | Medical specialist — trained on radiology, dermatology, ophthalmology |
| 4 | DeepSeek Reasoner | DeepSeek API | Clinical Council — chain-of-thought reasoning over findings (text-only, no images) |
Multi-model parallel analysis with cross-consensus
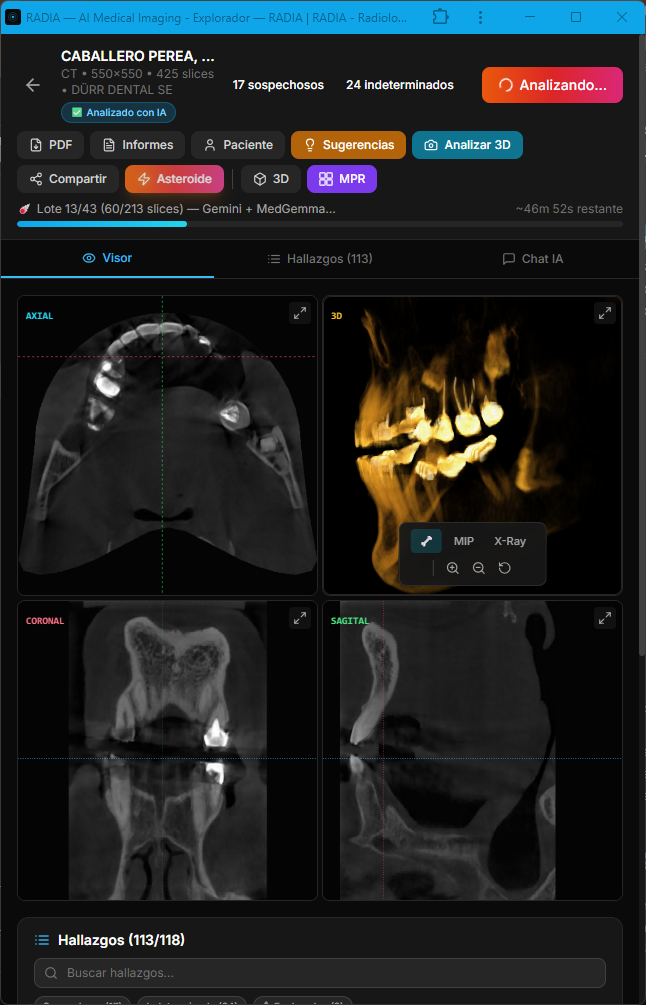
Asteroid Mode is the heavy artillery. Instead of using a single model that falls back if it fails, it runs Gemini and MedGemma in parallel on each slice, cross-references findings and marks matches as multi-model consensus. After visual detection, DeepSeek Reasoner acts as a "Clinical Council": it receives all findings (text only, no images) and applies chain-of-thought reasoning to debate clinical consistency, detect likely false positives, and prioritize findings.
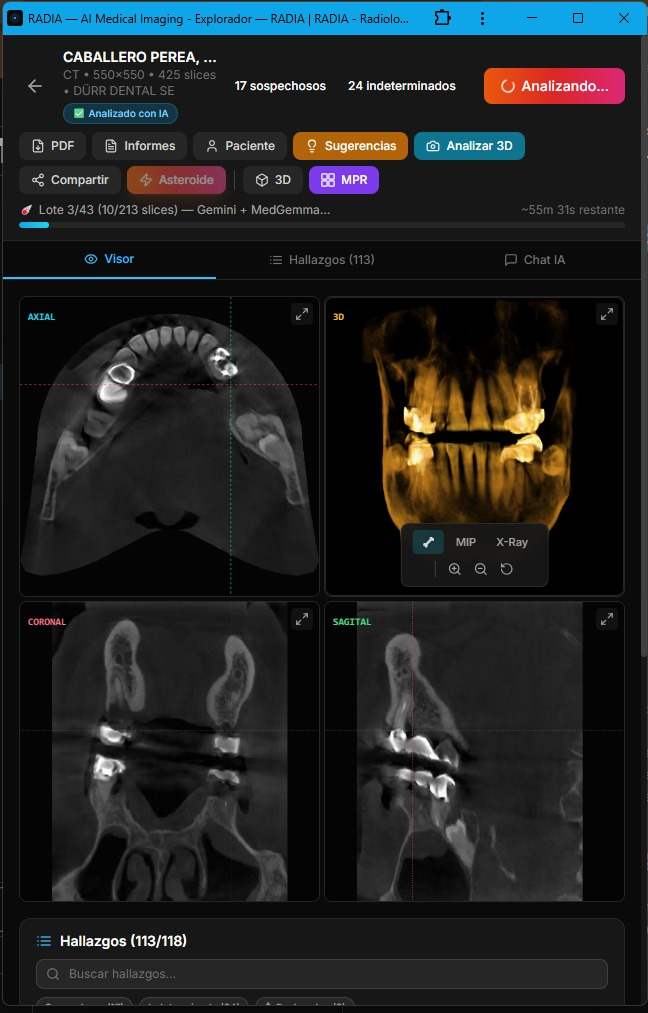
Asteroid Mode flow
Slice N of study
│
├─── Gemini 2.5 Flash ──────────┐
│ "3 findings detected" │
│ ├── Promise.allSettled()
├─── MedGemma 4B-IT ───────────┘
│ "2 findings detected"
│
▼
Cross-reference by (category + location + slice)
│
├── Match → ☄️ Consensus
│ • confidence × 1.15
│ • ai_model = "gemini-2.5-flash+medgemma-4b-it"
│
└── Single model only → normal finding
• confidence unchanged
• ai_model = individual model
▼
DeepSeek Reasoner — Clinical Council (text-only)
│
Input: all findings JSON + clinical data
│
Output:
├── Expanded clinical differential with reasoning
├── Likely false positive detection
├── Cross-finding correlations
└── Prioritization and recommendations| Aspect | Normal Scan | Asteroid Mode ☄️ |
|---|---|---|
| Sampling | Every 8 slices | Every 2 slices |
| Models | 1 (with fallback) | 2 in parallel |
| Deep analysis | Manual | Auto on top 5 findings |
| Time (~280 slices) | ~30 seconds | ~2-3 minutes |
| API cost | 1× | ~4× |
An important detail: AI consensus is never marked as doctor confirmation. The user_confirmed field always starts at 0 regardless of how many models agree. Human validation is a separate, intentional step — AI assists, it doesn't replace.
Anatomy of a finding
Each AI-generated finding is a rich structure with enough data for precise image localization, clinical categorization and validation workflow tracking. Findings on adjacent slices with the same category and location are automatically grouped (deduplicated), showing a slice range instead of repeated entries. The doctor can delete, hide from report, or navigate directly to each finding — individually or as a group.
Localization
- 📍 slice_number — slice where it was detected
- 📐 bounding_box — [x, y, w, h] normalized 0-1
- 🦷 location — anatomical text ("lower right molar area")
Classification
- 🏷️ category — periapical, fracture, mass, cyst...
- 🎯 confidence — 0.0 to 1.0
- 📏 measurements — dimensions in mm if applicable
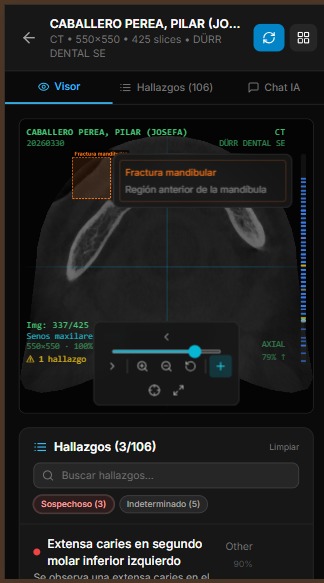
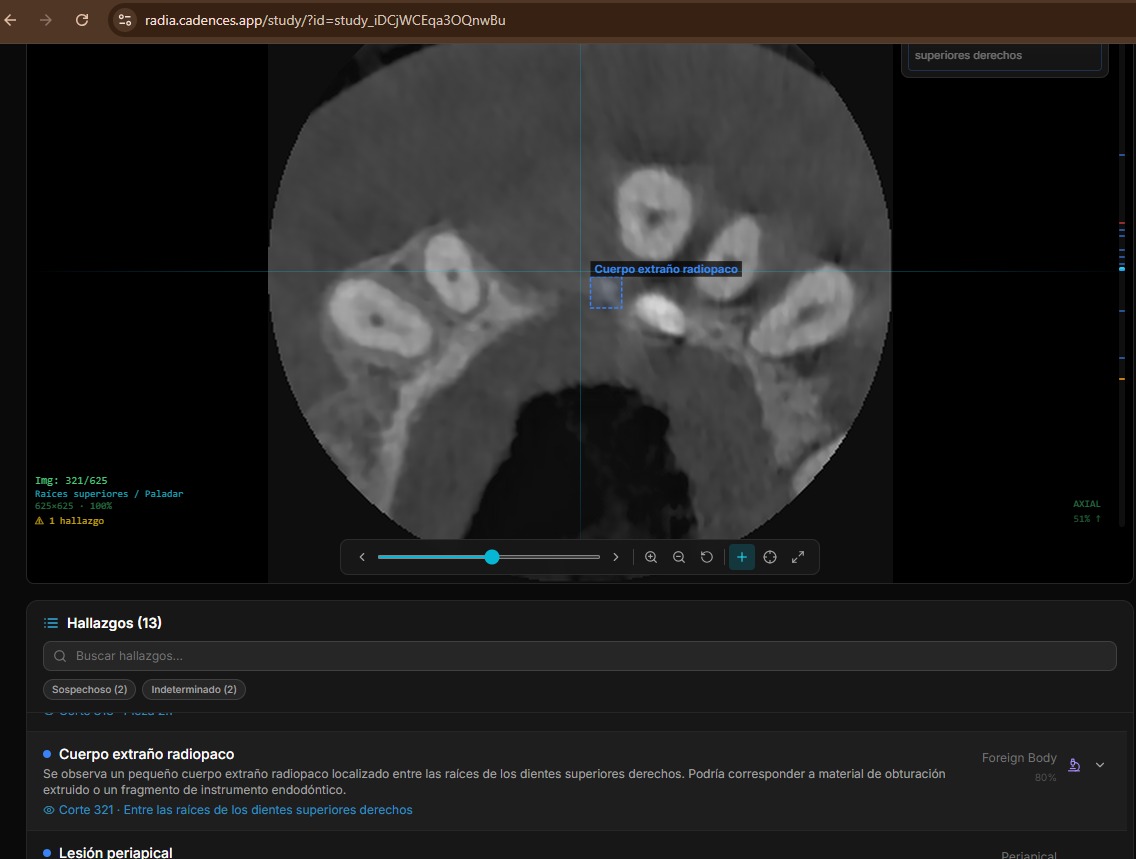
Severity is classified into 5 levels with consistent color coding across the entire UI:
Interactive markers on the volume
Findings with bounding boxes and slice numbers are projected as 3D markers onto the rendered volume. The bounding box (x, y) position maps to the volume coordinate system, slice_number determines Z depth, and the MVP (Model-View-Projection) matrix transforms them to screen coordinates every frame.
Tapping a marker shows a popup with finding information — title, severity, description, location — and three actions: go to findings list, jump to the 2D slice where the finding is, or open the MPR view centered on that position. The doctor navigates from 3D to 2D with a single touch. Additionally, Navigate Mode allows clicking anywhere on the 2D viewer to automatically jump to the nearest finding by Euclidean distance — ideal for exploring areas of interest without searching the list.
Technical detail: 3D → 2D projection
Markers are HTML divs positioned with position: absolute over the WebGL canvas. On each render frame, we multiply the finding's 3D coordinates by the MVP matrix and convert NDC to CSS percentages. If the marker ends up behind the camera (clip.w ≤ 0) or outside the frustum, it's hidden with opacity: 0.
16 study types, automatic detection
RADIA isn't just a dental viewer. It supports 16 study types organized in 6 medical sectors, each with specialized AI prompts and specific finding categories:
| Sector | Types | 3D/MPR |
|---|---|---|
| 🦷 Dental | CBCT, Panoramic, Periapical | CBCT ✓ |
| 🫁 Radiology | Chest X-Ray, Bone X-Ray, CT, Mammography | CT ✓ |
| 🧠 MRI | Brain, Musculoskeletal | Both ✓ |
| 📡 Ultrasound | General | — |
| 📷 Clinical Photo | Dermatology, Podiatry, Ophthalmology, Wounds, General | — |
| 🐾 Veterinary | General | — |
Detection is automatic: we analyze DICOM headers (Modality, SeriesDescription, StudyDescription) with a cascade of rules — first keywords in the description, then the DICOM modality code, then anatomy heuristics. The user can always correct manually, but in practice automatic detection is accurate 95%+ of the time.
Labrador with gastric dilatation-volvulus (GDV): from radiograph to signed report
The veterinary module isn't an afterthought bolted onto the medical engine — it's a dedicated branch with prompts tuned for animal anatomy, vet-specific clinical terminology, and a workflow built for 24/7 emergency cases. The AI distinguishes between laterolateral and ventrodorsal projections, recognizes the pathognomonic "double bubble" or "smurf hat" pattern, and prioritizes findings by life threat. In this real case, an abdominal radiograph of a Labrador Retriever triggered 19 automatic findings in under 30 seconds, with 6 classified as malignant.
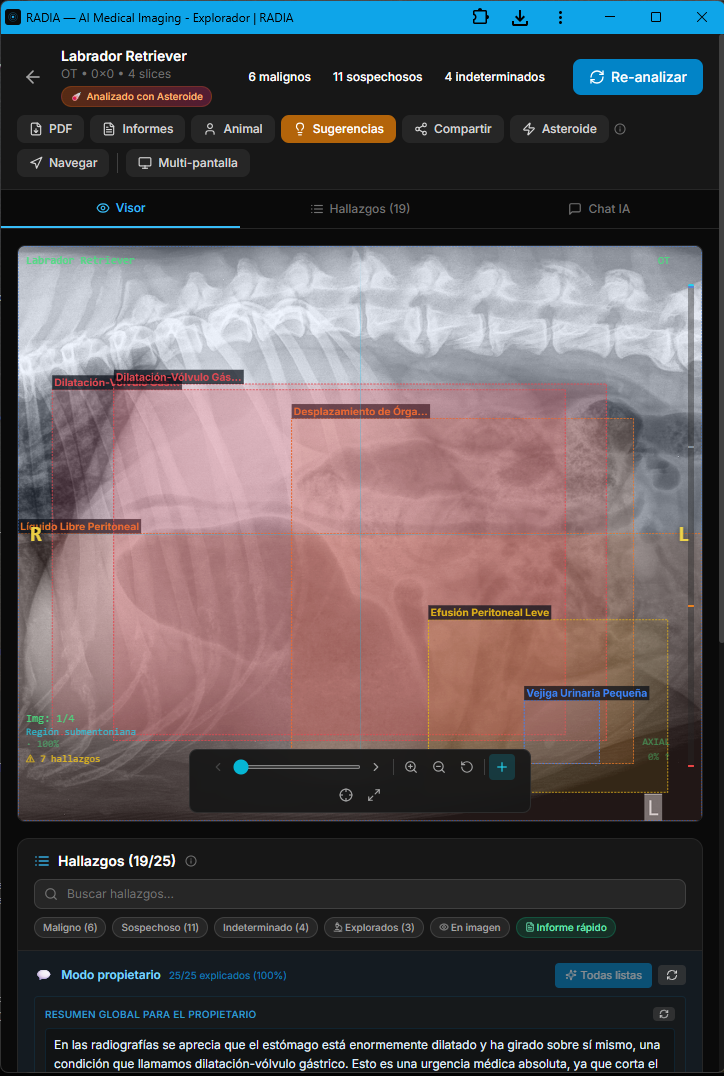
Asteroid Mode activates the full ensemble of models in parallel and applies a second critical-review pass — designed for emergency cases where missing a finding is not an option. The image also shows the semantic boxes: each color encodes severity (red=malignant, orange=suspicious, yellow=undetermined, green=normal) and each box is linked to a navigable finding in the bottom panel.
Owner mode: the AI explains the case without losing rigor
In veterinary medicine the "patient" doesn't talk — and the owner is the one signing the clinical decision. That's why RADIA generates, alongside the technical report, a plain-language narrative aimed at the animal's owner. Same clinical content, two registers: the technical one for the medical record, and the owner-facing one explaining what's visible, why it matters and how urgent it is. All traceable: every owner explanation points back to the technical finding that backs it.
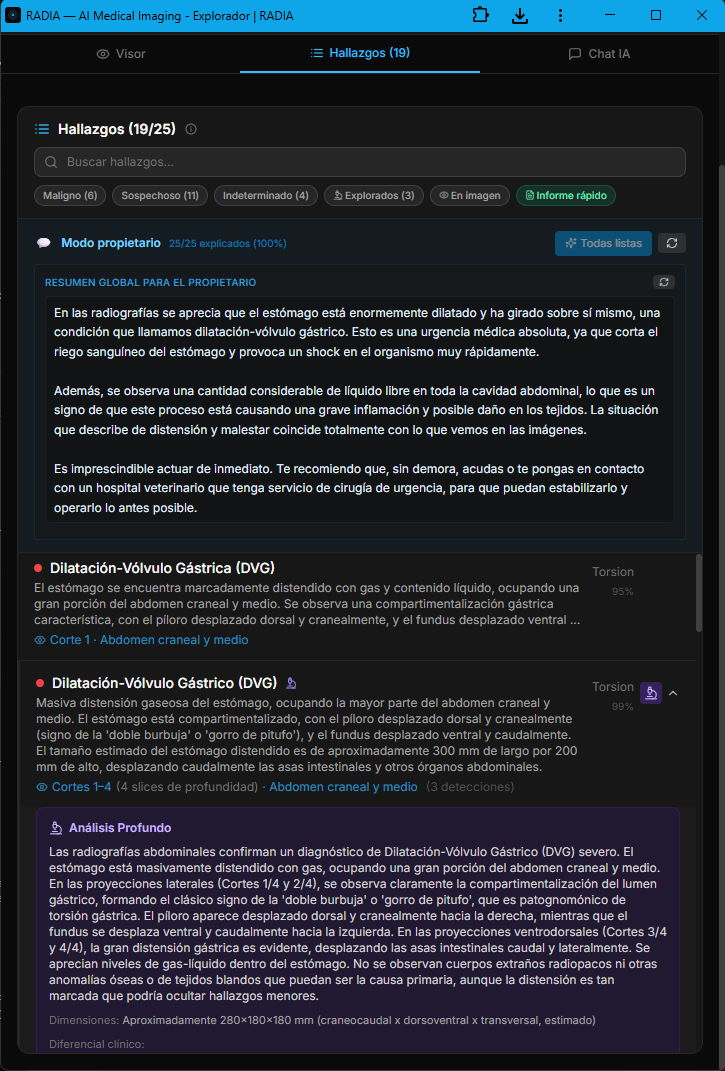
Appendix · numbered map of findings over the original image
The PDF the veterinarian downloads isn't a screenshot — it's a structured report generated client-side with jsPDF, signed with the professional's data (multi-country license number: Spain, Mexico, Colombia, Argentina, Chile, etc.), optional scanned stamp and clinic tax ID. The final page is always the cartographic appendix: a panoramic view with all findings numbered over the original radiograph, mirroring the severity colors and linking each number back to the detailed finding from earlier pages.
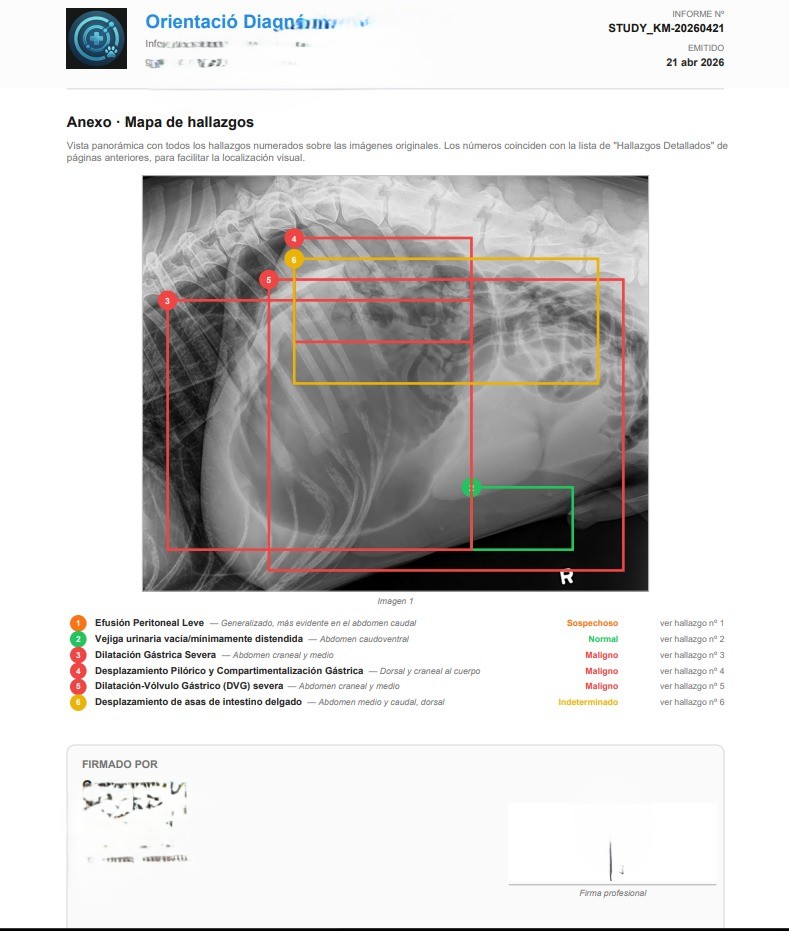
That closes the loop: AI detects, the professional validates and edits, the system writes the owner-facing narrative, and the signed PDF goes to the medical record. In this GDV case, RADIA turned a suspicious radiograph into an actionable report in under two minutes — the kind of timing that matters when you're dealing with an emergency whose therapeutic window is measured in hours.
Multimodal conversation with radiological context
The chat isn't a generic chatbot you pass an image to. It's an assistant with full study context: DICOM metadata, all previous findings, conversation history, and the current viewer image (including the 3D view if active). If the doctor asks "what do you think about the periapical area of tooth 46?", the AI knows which slice is shown, what findings were already detected there, and can generate a contextualized analysis.
If the 3D viewer is active, the chat automatically captures the current view as PNG and includes it in the request. The doctor rotates the volume to the perspective they're interested in, types their question, and the AI sees exactly what the doctor sees. No manual screenshots.
Share a study in one click
The doctor generates a secure link with optional expiration. The link provides read-only access to the complete study — DICOM viewer, 3D, MPR, all findings — without the recipient needing an account or login. For doctor-to-doctor collaboration (where authentication is needed), there's a "collab" mode that requires Google login but doesn't need study ownership.
The DICOM proxy verifies the sharing token against the database on every request. There are no signed URLs that could leak — if you revoke the link, access is cut immediately.
Two monitors, one synchronized study
Radiologists work with two or more monitors: one to review the image, another to consult the report, findings and history. Desktop PACS viewers support this natively, but web apps don't — opening two tabs of the same study gives two independent instances with no connection between them.
RADIA solves this with real-time synchronization between tabs. One click on "Multi-screen" opens a second window in viewer mode — optimized for the image, 3D and MPR — while the original becomes the editor with the findings list, AI chat and analysis tools. Everything the doctor does in one window is instantly reflected in the other.
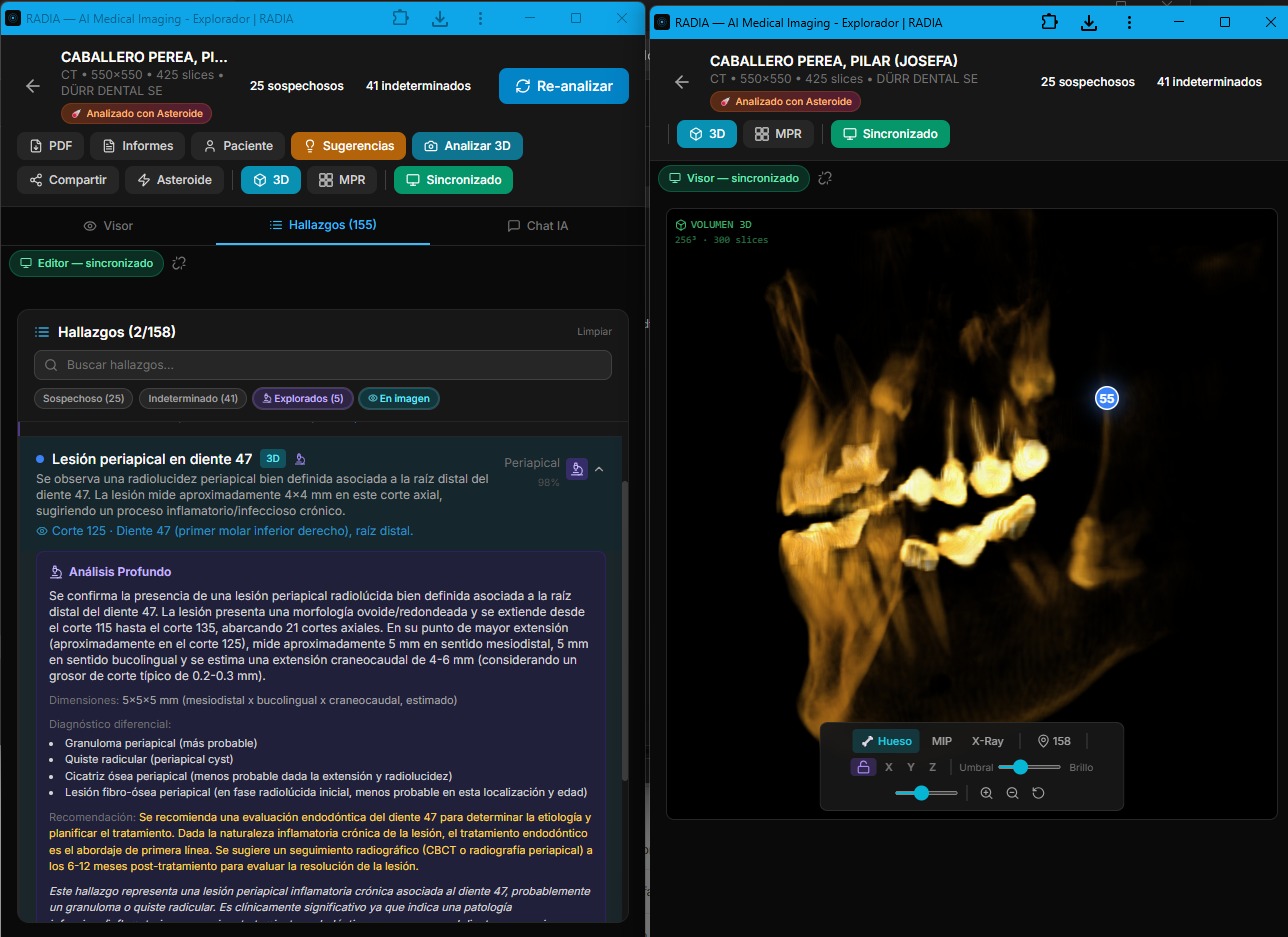
Real multi-monitor: deep analysis with clinical differential in editor (left) and 3D volume with synced markers in viewer (right).
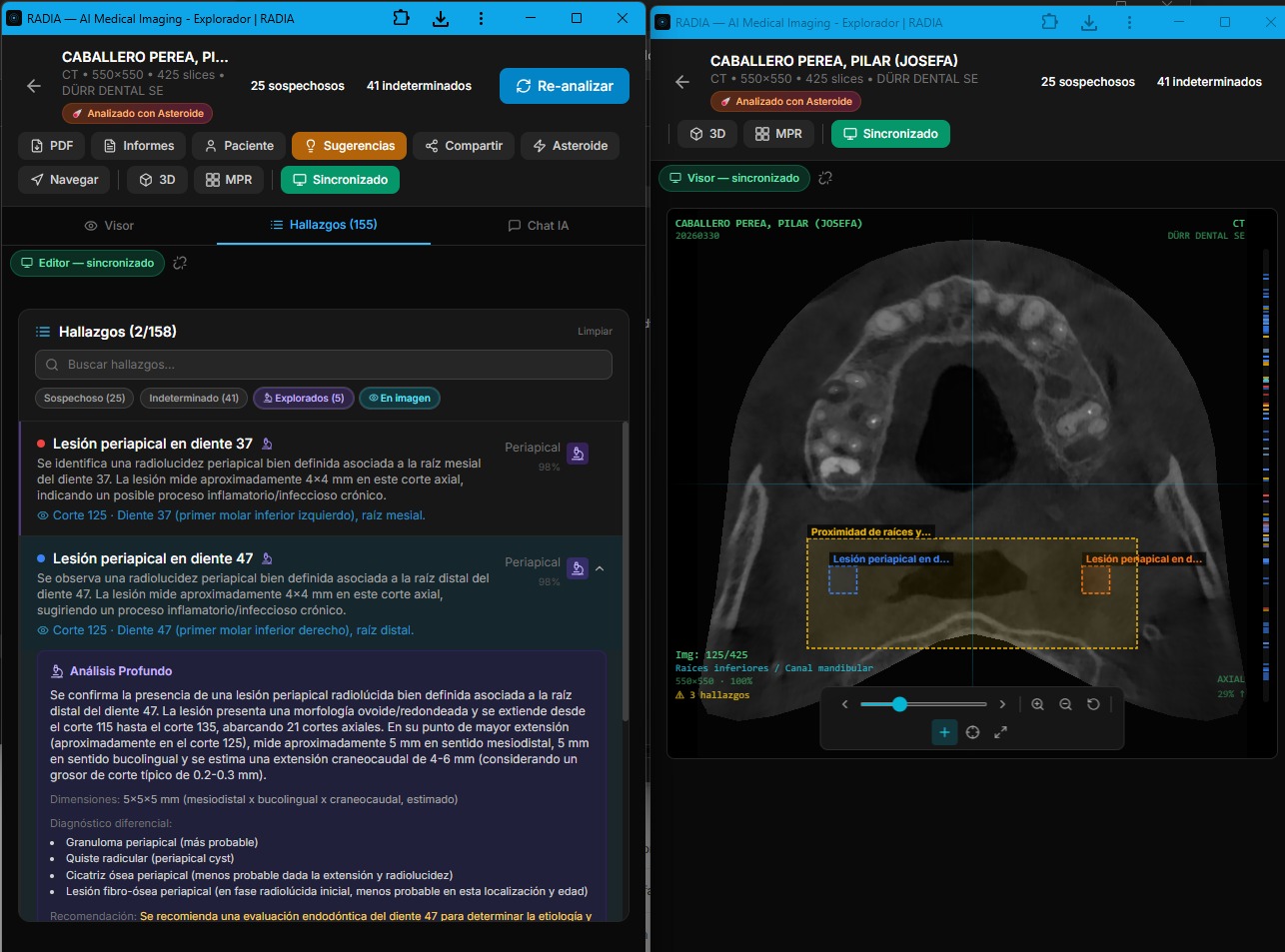
Synchronized 2D viewer: bounding boxes on the CBCT slice with labeled findings.
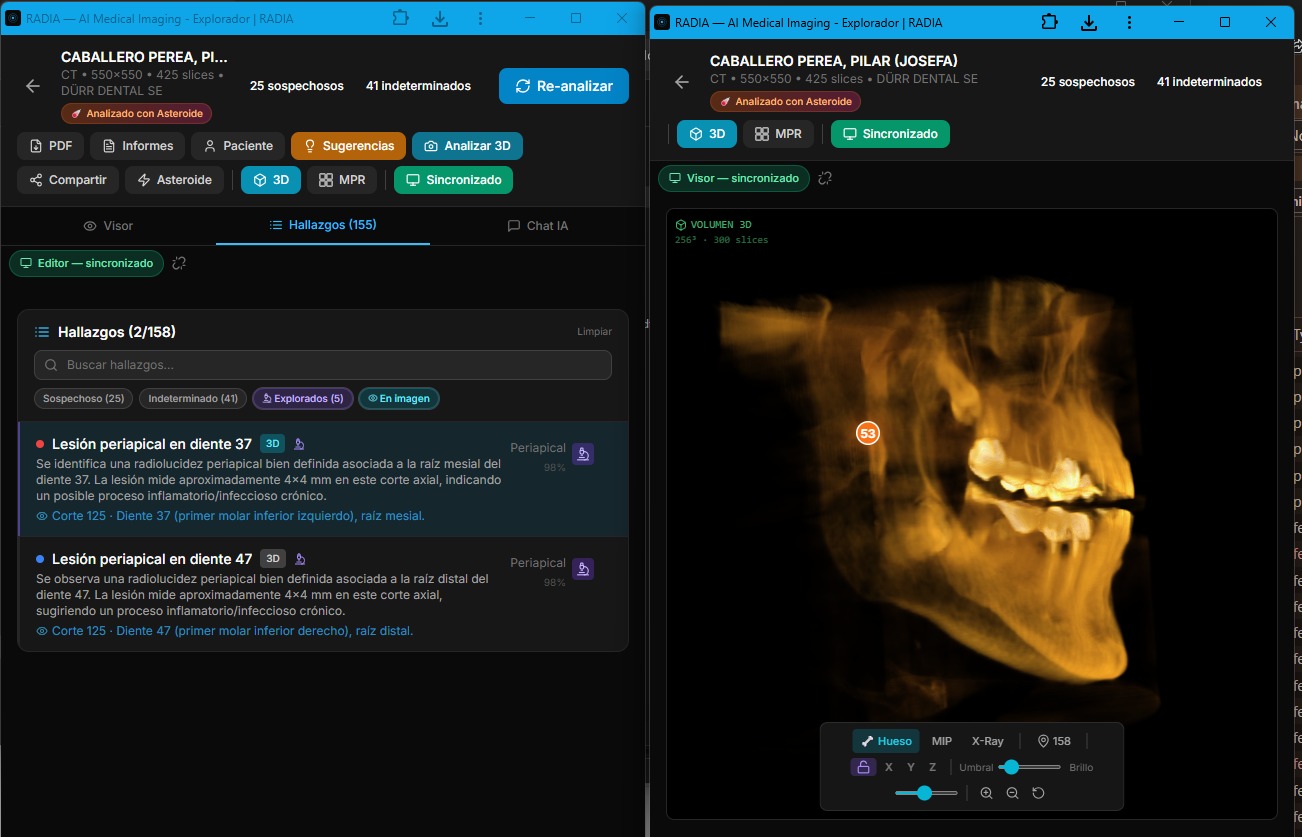
Findings list with severity filters and 3D with location markers.
BroadcastChannel: sync without a server
Synchronization uses the browser's BroadcastChannel API — a message channel between tabs of the same origin. No server involved, no WebSockets, no network latency. Message sent = message received in <1ms. Works on Chrome, Firefox, Edge and Safari — including installed PWAs.
What gets synchronized?
- 🔄 Slice navigation — scroll in one window, both change
- 🎯 Finding selection — click finding → viewer jumps to slice
- 🧊 3D/MPR view — switching visualization mode synced
- 💡 3D highlight — marking a finding on the volume
- 📊 Scan status — analysis progress visible in both
- 📝 New findings — on scan or deep analysis completion
- 🗑️ Deletions — deleting a finding updates both windows
- 💬 Chat findings — if AI saves a finding from chat
Sync protocol
Editor (monitor 1) Viewer (monitor 2)
│ │
│── NAVIGATE_SLICE { slice: 147 } ─────→│ Jumps to slice 147
│ │
│── SELECT_FINDING { id: "abc" } ──────→│ Shows bounding box
│ │
│── VIEW_MODE { mode: "3d" } ──────────→│ Activates 3D view
│ │
│── HIGHLIGHT_3D { id: "abc" } ────────→│ Marks on volume
│ │
│←── PING ──────────────────────────────│
│── PONG ──────────────────────────────→│ Heartbeat every 3s
│ │
│── FINDINGS_CHANGED ──────────────────→│ Reloads findingsA PING/PONG heartbeat every 3 seconds detects if the other window is still active. If it doesn't respond within 6 seconds, an amber disconnection indicator appears. On reconnection, it turns green. The doctor always knows if their windows are synchronized.
Editor Window
Findings list, AI chat, toolbar, analysis buttons, suggestions panel. The radiologist's control center.
Viewer Window
Full-screen image with 2D, 3D and MPR viewers. No distractions — just the image with bounding boxes and markers. Maximized on the second monitor.
Coming soon: cross-device sync
Phase 2 will take this synchronization beyond the same browser. Using Cloudflare Durable Objects + WebSocket, two different devices — e.g. the office PC and an iPad — will be able to sync in real time on the same study. Same message architecture, but with network transport instead of BroadcastChannel.
Client-generated radiological PDF
The PDF report is generated entirely in the browser with jsPDF — patient data, selected findings with severity codes, overall impression, recommendations and doctor's notes. The Worker never generates PDFs; the client has all the necessary information. This avoids the complexity of rendering documents in a serverless environment with 128MB of memory.
What we learned building this
Workers have a 30s timeout
You can't analyze 280 slices in a single request. The solution: the frontend orchestrates batches with a loop of POSTs. Each batch processes 8 slices, the Worker responds in ~10s, and the frontend shows real-time progress.
JPEG 2000 needs WASM
Half of medical DICOMs use JPEG 2000, which no browser decodes natively. OpenJPEG compiled to WASM solves this without server dependency, but weighs ~200KB. We load it lazily only when we detect Transfer Syntax 1.2.840.10008.1.2.4.90.
D1 is surprisingly fast
Distributed SQLite on the edge. Queries are ~1-3ms. For an app with dozens of inserts per analysis and multiple joins to fetch findings, D1 doesn't break a sweat. The real limitation is concurrent writes, which we avoid with the sequential batch pattern.
WebGL2 is enough for this
You don't need Three.js for medical volume rendering. Two shaders, native WebGL2 3D textures, and ~300 lines of 4×4 matrix math. The result: 0 3D dependencies, lightweight bundle, and full control of the rendering pipeline.
Medical AI gets it wrong (a lot)
Gemini and MedGemma are impressive but not infallible. That's why the confirmation flow is explicit: the doctor reviews each finding and marks it as confirmed or rejected. Asteroid Mode reduces errors with multi-model consensus, but never claims to replace the professional.
R2 for medical files scales perfectly
A dental CBCT is ~150MB of DICOMs. R2 serves them by prefix with paginated list() and batch delete(). No egress fees, no cold starts. The key: separate original DICOMs (for AI) from PNG thumbnails (for fast navigation).
A medical viewer that shouldn't be possible on the edge
RADIA proves that professional medical tools can live entirely on the edge, with no dedicated servers, no rented GPUs, no infrastructure to maintain. A doctor opens a browser, uploads their study, and three AI models analyze every slice while the 3D viewer rebuilds in WebGL2 at 30fps.
The full stack — Astro, React, Cloudflare Workers, D1, R2, Gemini 2.5 Flash, MedGemma, WebGL2 — deploys with npm run deploy in 15 seconds. No containers, no CI/CD pipelines, no clusters. The future of medical tooling is a static build with serverless edge functions, and it's already here.
Full stack
🩻
RADIA is part of Cadences
One platform, multiple storefronts. RADIA shares infrastructure (D1, auth, billing) with the rest of the Cadences ecosystem. Each storefront is independent in code but unified in data.